Mating of yeast
The mating of yeast, also known as yeast sexual reproduction, is a biological process that promotes genetic diversity and adaptation in yeast species. Yeast species, such as Saccharomyces cerevisiae (baker's yeast), are single-celled eukaryotes that can exist as either haploid cells, which contain a single set of chromosomes, or diploid cells, which contain two sets of chromosomes. Haploid yeast cells come in two mating types, a and α, each producing specific pheromones to identify and interact with the opposite type, thus displaying simple sexual differentiation.[a] A yeast cell's mating type is determined by a specific genetic locus known as MAT, which governs its mating behaviour. Haploid yeast can switch mating types through a form of genetic recombination, allowing them to change mating type as often as every cell cycle. When two haploid cells of opposite mating types encounter each other, they undergo a complex signaling process that leads to cell fusion and the formation of a diploid cell. Diploid cells can reproduce asexually, but under nutrient-limiting conditions, they undergo meiosis to produce new haploid spores.
The differences between a and α cells, driven by specific gene expression patterns regulated by the MAT locus, are crucial for the mating process. Additionally, the decision to mate involves a highly sensitive and complex signaling pathway that includes pheromone detection and response mechanisms. In nature, yeast mating often occurs between closely related cells, although mating type switching and pheromone signaling allow for occasional outcrossing to enhance genetic variation. Furthermore, certain yeast species, like Schizosaccharomyces pombe and Cryptococcus neoformans, have unique mating behaviours and regulatory mechanisms, demonstrating the diversity and adaptability of yeast reproductive strategies.
Mating types
[edit]Yeast cells can stably exist in either a diploid or a haploid form. Both haploid and diploid yeast cells reproduce by mitosis, in which daughter cells bud from mother cells. Haploid cells are capable of mating with other haploid cells of the opposite mating type (an a cell can only mate with an α cell and vice versa) to produce a stable diploid cell. Diploid cells, usually upon facing stressful conditions like nutrient depletion, can undergo meiosis to produce four haploid spores: two a spores and two α spores.[1][2]
Differences between a and α cells
[edit]
a cells produce a-factor, a mating pheromone which signals the presence of an a cell to neighbouring α cells.[3] a cells respond to α-factor, the α cell mating pheromone, by growing a projection (known as a shmoo, due to its distinctive shape resembling the Al Capp cartoon character Shmoo) towards the source of α-factor.[4] Similarly, α cells produce α-factor, and respond to a-factor by growing a projection towards the source of the pheromone.[5] The selective response of haploid cells to the mating pheromones of the opposite mating type allows mating between a and α cells, but not between cells of the same mating type.[6]
These phenotypic differences between a and α cells are due to a different set of genes being actively transcribed and repressed in cells of the two mating types. a cells activate genes which produce a-factor and produce a cell surface receptor (Ste2) which binds to α-factor and triggers signaling within the cell.[7][8] a cells also repress the genes associated with being an α cell. Conversely, α cells activate genes which produce α-factor and produce a cell surface receptor (Ste3) which binds and responds to a-factor, and α cells repress the genes associated with being an a cell.[9]
MAT locus
[edit]The different sets of transcriptional repression and activation, which characterize a and α cells, are caused by the presence of one of two alleles for a mating-type locus called MAT: MATa or MATα located on chromosome III.[10] The MAT locus is usually divided into five regions (W, X, Y, Z1, and Z2) based on the sequences shared among the two mating types.[11] The difference lie in the Y region (Ya and Yα), which contains most of the genes and promoters.[7]
The MATa allele of MAT encodes a gene called a1, which directs the a-specific transcriptional program (such as expressing STE2 and repressing STE3) that defines an a haploid cell. The MATα allele of MAT encodes the α1 and α2 genes, which directs the α-specific transcriptional program (such as expressing STE3, repressing STE2, and producing prepro-α-factor) that defines an α haploid cell.[7] S. cerevisiae has an a2 gene with no apparent function that shares much of its sequence with α2; however, other yeast species like Candida albicans do have a functional and distinct MATa2 gene.[6][10]
Differences between haploid and diploid cells
[edit]
Haploid cells are one of two mating types (a or α) and respond to the mating pheromone produced by haploid cells of the opposite mating type.[4] Haploid cells cannot undergo meiosis.[12] Diploid cells do not produce or respond to either mating pheromone and do not mate, but they can undergo meiosis to produce four haploid cells.[13]
Like the differences between haploid a and α cells, different patterns of gene repression and activation are responsible for the phenotypic differences between haploid and diploid cells.[14] In addition to the transcriptional patterns of a and α cells, haploid cells of both mating types share a haploid transcriptional pattern which activates haploid-specific genes (such as HO) and represses diploid-specific genes (such as IME1).[15] Conversely, diploid cells activate diploid-specific genes and repress haploid-specific genes.[16]
The different gene expression patterns of haploid and diploid cells are attributable to the MAT locus. Haploid cells only contain one copy of each of the 16 chromosomes and therefore only possess one MAT allele (either MATa or MATα), which determines their mating type.[17] Diploid cells result from the mating of an a cell and an α cell, and they possess 32 chromosomes (in 16 pairs), including one chromosome bearing the MATa allele and another chromosome bearing the MATα allele.[18] The combination of the information encoded by the MATa allele (the a1 gene) and the MATα allele (the α1 and α2 genes) triggers the diploid transcriptional program.[19] Conversely, the presence of only one MAT allele, either MATa or MATα, triggers the haploid transcriptional program.[20][7]
Through genetic engineering, a MATa allele can be added to a MATα haploid cell, causing it to behave like a diploid cell.[21] The cell will not produce or respond to mating pheromones, and when starved, the cell will unsuccessfully attempt to undergo meiosis with fatal results.[22] Similarly, deletion of one copy of the MAT locus in a diploid cell, leaving either a MATa or MATα allele, will cause a diploid cell to behave like a haploid cell of the associated mating type.[23][24]
a-like faker cells
[edit]α cells with inactivated α1 and α2 genes at the MAT locus will exhibit the mating behavior of a cells. When an a-like faker (alf) cell mates with an α cell, they form a diploid cell lacking an active copy of the a1 gene. As a result, these diploid cells cannot form the a1-α2 protein complex needed to repress haploid-specific genes. This diploid cell will act like a haploid α cell, producing α pheromones to mate with an a haploid cell, resulting in aneuploidy.[25]
Since α cells do not ordinarily mate with each other, the presence of a-like faker cells in a population of α cells can be detected in an a-like faker assay. This test exposes the MATα population, which lacks an active copy of the HIS3 gene, to a tester strain like YPH316 yeast, which lack a HIS1 gene, on YEPD agar. After transferring the pairs of yeast strains onto Sabouraud agar, only those that formed diploid cells by having a-like faker cells mate with the tester strain will be capable of synthesizing the amino acid histidine to survive. The extent of chromosome instability can be inferred from the proportion of surviving pairs since a-like faker cells naturally arise from damage to Chromosome III in yeast cells.[26]
Decision to mate
[edit]Mating in yeast is stimulated by a cells' a-factor or α cells' α-factor pheromones binding the Ste3 receptor of α cells or Ste2 receptor of a cells, respectively, activating a heterotrimeric G protein.[27][28][29] The dimeric portion of this G-protein recruits Ste5 and its MAPK cascade to the membrane, resulting in the phosphorylation of Fus3.[30]
The switching mechanism arises as a result of competition between the Fus3 protein (a MAPK protein) and the phosphatase Ptc1.[31] These proteins both attempt to control the four phosphorylation sites of Ste5, a scaffold protein, with Fus3 attempting to phosphorylate the phosphosites and Ptc1 attempting to dephosphorylate them.[32]
Presence of α-factor induces recruitment of Ptc1 to Ste5 via a four-amino acid motif located within the Ste5 phosphosites.[33] Ptc1 then dephosphorylates Ste5, resulting in the dissociation of the Fus3-Ste5 complex.[34] Fus3 dissociates in a switch-like manner, dependent on the phosphorylation state of the four phosphosites.[35] All four phosphosites must be dephosphorylated in order for Fus3 to dissociate.[36][37] Fus3's ability to compete with Ptc1 decreases as Ptc1 is recruited, and thus the rate of dephosphorylation increases with the presence of pheromone.[38]
Kss1, a homologue of Fus3, does not affect shmooing, and does not contribute to the switch-like mating decision.[39][40]
In yeast, mating as well as the production of shmoos occur via an all-or-none, switch-like mechanism.[41] This switch-like mechanism allows yeast cells to avoid making an unwise commitment to a highly demanding procedure.[42] The decision to mate must balance being energy-conservative and fast enough to avoid losing the potential mate.[43]
Yeast maintain an ultra-sensitivity to mating through:
- Multi-site phosphorylation – Fus3 only dissociates from Ste5 and becomes fully active when all four of the phosphosites are dephosphorylated. Even one phosphorylated site will result in immunity to α-factor.[44]
- Two-stage binding – Fus3 and Ptc1 bind to separate docking sites on Ste5. Only after docking can they act on the phosphosites.[45]
- Steric hindrance – competition between Fus3 and Ptc1 to control the four phosphosites on Ste3
a and α yeast share the same mating response pathway, with the only difference being the type of receptor that each mating type possesses.[46] Thus, the above description of an a-type yeast stimulated with α-factor resembles the mechanism of an α-type yeast stimulated with a-factor.[47][48]
Mating type switching
[edit]
Wild type haploid yeast are capable of switching mating type between a and α.[49] Consequently, even if a single haploid cell of a given mating type founds a colony of yeast, mating type switching will cause cells of both a and α mating types to be present in the population.[50][51] Combined with the strong drive for haploid cells to mate with cells of the opposite mating type and form diploids, mating type switching and consequent mating will cause the majority of cells in a colony to be diploid, regardless of whether a haploid or diploid cell founded the colony.[52] The vast majority of yeast strains studied in laboratories have been altered such that they cannot perform mating type switching (by deletion of the HO gene; see below). This allows the stable propagation of haploid yeast, as haploid cells of the a mating type will remain a cells (and α cells will remain α cells), unable to form diploid cells unless artificially exposed to the other mating type.[53]

HML and HMR: the silent mating cassettes
[edit]Haploid yeast switch mating type by replacing the information present at the MAT locus.[54] For example, an a cell will switch to an α cell by replacing the MATa allele with the MATα allele.[55] This replacement of one allele of MAT for the other is possible because yeast cells carry an additional silenced copy of both the MATa and MATα alleles: the HML (homothallic mating left) locus typically carries a silenced copy of the MATα allele, and the HMR (homothallic mating right) locus typically carries a silenced copy of the MATa allele.[7] The silent HML and HMR loci are often referred to as the silent mating cassettes, as the information present there is 'read into' the active MAT locus.[56]
These additional copies of the mating type information do not interfere with the function of whatever allele is present at the MAT locus because they are not expressed, so a haploid cell with the MATa allele present at the active MAT locus is still an a cell, despite also having a silenced copy of the MATα allele present at HML.[57] Only the allele present at the active MAT locus is transcribed, and thus only the allele present at MAT will influence cell behaviour.[6] Hidden mating type loci are epigenetically silenced by SIR proteins, which form a heterochromatin scaffold that prevents transcription from the silent mating cassettes.[58]
Mechanics of the mating type switch
[edit]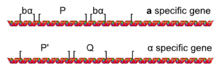
The process of mating type switching is a gene conversion event initiated by the HO gene.[59] The HO gene is a tightly regulated haploid-specific gene that is only activated in haploid cells during the G1 phase of the cell cycle.[60] The protein encoded by the HO gene is a DNA endonuclease, which physically cleaves DNA, but only at the MAT locus (due to the DNA sequence specificity of the HO endonuclease).[61]
Once HO cuts the DNA at MAT, exonucleases are attracted to the cut DNA ends and begin to degrade the DNA on both sides of the cut site.[62] This DNA degradation by exonucleases eliminates the DNA which encoded the MAT allele; however, the resulting gap in the DNA is repaired by copying in the genetic information present at either HML or HMR, filling in a new allele of either the MATa or MATα gene. Thus, the silenced alleles of MATa and MATα present at HML and HMR serve as a source of genetic information to repair the HO-induced DNA damage at the active MAT locus.[7]
Directionality of the mating type switch
[edit]The repair of the MAT locus after cutting by the HO endonuclease almost always results in a mating type switch.[7][61] When an a cell cuts the MATa allele present at the MAT locus, the cut at MAT will almost always be repaired by copying the information present at HML.[6] This results in MAT being repaired to the MATα allele, switching the mating type of the cell from a to α.[63] Similarly, an α cell which has its MATα allele cut by the HO endonuclease will almost always repair the damage using the information present at HMR, copying the MATa gene to the MAT locus and switching the mating type of α cell to a.[64]
This is the result of a recombination enhancer (RE) located on the left arm of chromosome III.[65] Normally, a cells have Mcm1 bind to the RE to promote recombination using the HML region.[66] Deletion of the RE causes a cells to instead repair using HMR, maintaining their status as a cells rather than switching mating types.[67] In α cells, the α2 factor binds at the RE to repress recombination using the HML region.[68] Thus, yeast have an innate bias toward DNA repair of the MAT locus using the HMR region.[69]
Mating and inbreeding
[edit]In 2006, evolutionary geneticist Leonid Kruglyak found that S. cerevisiae matings only involve out-crossing between different strains roughly once every 50,000 cell divisions. The vast majority of yeast mating instead involves members of the same strain because mating type switching allows a single ascus to produce both mating types from a single haploid cell.[70] This suggests that yeast primarily maintain their capability to mate through recombinational DNA repair during meiosis, rather than natural selection for fitness among a population with high genetic variability.[71]
Special cases
[edit]Fission yeast
[edit]Schizosaccharomyces pombe is a facultative sexual yeast that can undergo mating when nutrients are limited.[72] Exposure of S. pombe to hydrogen peroxide, which causes oxidative stress to DNA, strongly induces mating, meiosis, and formation of meiotic spores.[73] Thus, meiosis and meiotic recombination may be an adaptation for repairing DNA damage.[74] The MAT locus' structure in S. pombe resembles S. cerevisiae. The mating-type switching system is similar but evolved independently.[6]
Self-mating in Cryptococcus neoformans
[edit]Cryptococcus neoformans is a basidiomycetous fungus that grows as a budding yeast in culture and infected hosts. C. neoformans causes life-threatening meningoencephalitis in immunocompromised patients. It undergoes a filamentous transition during the sexual cycle to produce spores, the suspected infectious agent. The vast majority of environmental and clinical isolates of C. neoformans are of mating type α. Filaments ordinarily have haploid nuclei, but these can undergo a process of diploidization (perhaps by endoreduplication or stimulated nuclear fusion) to form diploid cells termed blastospores.[75]
The diploid nuclei of blastospores can then undergo meiosis, including recombination, to form haploid basidiospores that can then be dispersed.[75] This process is referred to as monokaryotic fruiting. This process depends on the gene dmc1, a conserved homologue of the bacterial RecA and eukaryotic RAD51 genes. Dmc1 mediates homologous chromosome pairing during meiosis and repair of double-strand breaks in DNA.[76] Meiosis in C. neoformans may be performed to promote DNA repair in DNA-damaging environments, such as host-mediated responses to infection.[75]
Notes
[edit]- ^ For the sake of clarity, this article bolds the Latin letter "a" and uses regular font weight for the Greek α. The usual convention is to print both in the same weight, but doing so would make the two letters hard to tell apart in italicized text.
References
[edit]- ^ Bergman LW (2001). "Growth and maintenance of yeast". Two-Hybrid Systems. Methods in Molecular Biology (Clifton, N.J.). Vol. 177. pp. 9–14. doi:10.1385/1-59259-210-4:009. ISBN 1-59259-210-4. PMID 11530618.
- ^ Börner GV, Hochwagen A, MacQueen AJ (October 2023). "Meiosis in budding yeast". Genetics. 225 (2). doi:10.1093/genetics/iyad125. PMC 10550323. PMID 37616582.
- ^ Michaelis S, Barrowman J (September 2012). "Biogenesis of the Saccharomyces cerevisiae pheromone a-factor, from yeast mating to human disease". Microbiology and Molecular Biology Reviews. 76 (3): 626–651. doi:10.1128/MMBR.00010-12. PMC 3429625. PMID 22933563.
- ^ a b Merlini L, Dudin O, Martin SG (March 2013). "Mate and fuse: how yeast cells do it". Open Biology. 3 (3): 130008. doi:10.1098/rsob.130008. PMC 3718343. PMID 23466674.
- ^ Chen W, Nie Q, Yi TM, Chou CS (July 2016). Edelstein-Keshet L (ed.). "Modelling of Yeast Mating Reveals Robustness Strategies for Cell-Cell Interactions". PLOS Computational Biology. 12 (7): e1004988. Bibcode:2016PLSCB..12E4988C. doi:10.1371/journal.pcbi.1004988. PMC 4942089. PMID 27404800.
- ^ a b c d e Hanson SJ, Wolfe KH (May 2017). "An Evolutionary Perspective on Yeast Mating-Type Switching". Genetics. 206 (1): 9–32. doi:10.1534/genetics.117.202036. PMC 5419495. PMID 28476860.
- ^ a b c d e f g Haber JE (May 2012). "Mating-type genes and MAT switching in Saccharomyces cerevisiae". Genetics. 191 (1): 33–64. doi:10.1534/genetics.111.134577. PMC 3338269. PMID 22555442.
- ^ Gastaldi S, Zamboni M, Bolasco G, Di Segni G, Tocchini-Valentini GP (August 2016). "Analysis of random PCR-originated mutants of the yeast Ste2 and Ste3 receptors". MicrobiologyOpen. 5 (4): 670–686. doi:10.1002/mbo3.361. PMC 4985600. PMID 27150158.
- ^ Brenner S, Miller JH (2001). "Cassette Model". Encyclopedia of Genetics. Academic Press. pp. 275–278. doi:10.1006/rwgn.2001.0162. ISBN 978-0-12-227080-2. Retrieved 2024-05-09.
- ^ a b Tsong AE, Miller MG, Raisner RM, Johnson AD (November 2003). "Evolution of a combinatorial transcriptional circuit: a case study in yeasts". Cell. 115 (4): 389–399. doi:10.1016/S0092-8674(03)00885-7. PMID 14622594.
- ^ Lee CS, Haber JE (April 2015). Gellert M, Craig N (eds.). "Mating-type Gene Switching in Saccharomyces cerevisiae". Microbiology Spectrum. 3 (2): MDNA3–0013–2014. doi:10.1128/microbiolspec.MDNA3-0013-2014. PMID 26104712.
- ^ Wagstaff JE, Klapholz S, Esposito RE (May 1982). "Meiosis in haploid yeast". Proceedings of the National Academy of Sciences of the United States of America. 79 (9): 2986–2990. Bibcode:1982PNAS...79.2986W. doi:10.1073/pnas.79.9.2986. PMC 346333. PMID 7045878.
- ^ Yeager R, Bushkin GG, Singer E, Fu R, Cooperman B, McMurray M (August 2021). "Post-Transcriptional Control of Mating-Type Gene Expression during Gametogenesis in Saccharomyces cerevisiae". Biomolecules. 11 (8): 1223. doi:10.3390/biom11081223. PMC 8394074. PMID 34439889.
- ^ Solieri L, Cassanelli S, Huff F, Barroso L, Branduardi P, Louis EJ, et al. (December 2021). "Insights on life cycle and cell identity regulatory circuits for unlocking genetic improvement in Zygosaccharomyces and Kluyveromyces yeasts". FEMS Yeast Research. 21 (8). doi:10.1093/femsyr/foab058. PMC 8673824. PMID 34791177.
- ^ Nagaraj VH, O'Flanagan RA, Bruning AR, Mathias JR, Vershon AK, Sengupta AM (August 2004). "Combined analysis of expression data and transcription factor binding sites in the yeast genome". BMC Genomics. 5 (1): 59. doi:10.1186/1471-2164-5-59. PMC 517709. PMID 15331021.
- ^ Voth WP, Stillman DJ (September 2003). "Changes in developmental state: demolish the old to construct the new". Genes & Development. 17 (18): 2201–2204. doi:10.1101/gad.1142103. PMID 12975315.
- ^ Watkinson SC, Boddy L, Money NP (2016). "Chapter 4 - Genetics – Variation, Sexuality, and Evolution". The Fungi (3rd ed.). Academic Press. pp. 99–139. doi:10.1016/B978-0-12-382034-1.00004-9. ISBN 978-0-12-382034-1.
- ^ Bizzarri M, Giudici P, Cassanelli S, Solieri L (2016-04-11). Fairhead C (ed.). "Chimeric Sex-Determining Chromosomal Regions and Dysregulation of Cell-Type Identity in a Sterile Zygosaccharomyces Allodiploid Yeast". PLOS ONE. 11 (4): e0152558. Bibcode:2016PLoSO..1152558B. doi:10.1371/journal.pone.0152558. PMC 4827841. PMID 27065237.
- ^ Zill OA, Rine J (June 2008). "Interspecies variation reveals a conserved repressor of alpha-specific genes in Saccharomyces yeasts". Genes & Development. 22 (12): 1704–1716. doi:10.1101/gad.1640008. PMC 2428066. PMID 18559484.
- ^ Leupold U (February 1980). "Transposable mating-type genes in yeasts". Nature. 283 (5750): 811–812. Bibcode:1980Natur.283..811L. doi:10.1038/283811a0. PMID 6987523.
- ^ Liu Y, Huang Y, Lu R, Xin F, Liu G (2022). "Synthetic biology applications of the yeast mating signal pathway". Trends in Biotechnology. 40 (5): 620–631. doi:10.1016/j.tibtech.2021.09.007. PMID 34666896.
- ^ Lengeler KB, Davidson RC, D'souza C, Harashima T, Shen WC, Wang P, et al. (December 2000). "Signal transduction cascades regulating fungal development and virulence". Microbiology and Molecular Biology Reviews. 64 (4): 746–785. doi:10.1128/MMBR.64.4.746-785.2000. PMC 99013. PMID 11104818.
- ^ Lee K, Neigeborn L, Kaufman RJ (April 2003). "The unfolded protein response is required for haploid tolerance in yeast". The Journal of Biological Chemistry. 278 (14): 11818–11827. doi:10.1074/jbc.M210475200. PMID 12560331.
- ^ Conn PM (2006). "17 - Telomeres and Aging in the Yeast Model System". Handbook of Models for Human Aging. Academic Press. pp. 191–205. doi:10.1016/B978-0-12-369391-4.X5000-0. ISBN 978-0-12-369391-4.
- ^ Fraser JA, Heitman J (23 February 2004). "Evolution of Fungal Sex Chromosomes". Molecular Microbiology. 51 (2): 299–306. doi:10.1046/j.1365-2958.2003.03874.x. ISSN 0950-382X.
- ^ Novoa CA, Ang JS, Stirling PC (17 October 2017). "The A-Like Faker Assay for Measuring Yeast Chromosome III Stability". In Muzi-Falconi M, Brown GW (eds.). Genome Instability. Springer Nature. pp. 1–9. ISBN 978-1-4939-7305-7.
- ^ Versele M, Lemaire K, Thevelein JM (July 2001). "Sex and sugar in yeast: two distinct GPCR systems". EMBO Reports. 2 (7): 574–579. doi:10.1093/embo-reports/kve132. PMC 1083946. PMID 11463740.
- ^ Emmerstorfer-Augustin A, Augustin CM, Shams S, Thorner J (November 2018). Glick BS (ed.). "Tracking yeast pheromone receptor Ste2 endocytosis using fluorogen-activating protein tagging". Molecular Biology of the Cell. 29 (22): 2720–2736. doi:10.1091/mbc.E18-07-0424. PMC 6249837. PMID 30207829.
- ^ Navarro-Olmos R, Kawasaki L, Domínguez-Ramírez L, Ongay-Larios L, Pérez-Molina R, Coria R (February 2010). Boone C (ed.). "The beta subunit of the heterotrimeric G protein triggers the Kluyveromyces lactis pheromone response pathway in the absence of the gamma subunit". Molecular Biology of the Cell. 21 (3): 489–498. doi:10.1091/mbc.e09-06-0472. PMC 2814793. PMID 20016006.
- ^ Winters MJ, Pryciak PM (April 2019). Boone C (ed.). "MAPK modulation of yeast pheromone signaling output and the role of phosphorylation sites in the scaffold protein Ste5". Molecular Biology of the Cell. 30 (8): 1037–1049. doi:10.1091/mbc.E18-12-0793. PMC 6589907. PMID 30726174.
- ^ Malleshaiah MK, Shahrezaei V, Swain PS, Michnick SW (May 2010). "The scaffold protein Ste5 directly controls a switch-like mating decision in yeast". Nature. 465 (7294): 101–105. Bibcode:2010Natur.465..101M. doi:10.1038/nature08946. PMID 20400943.
- ^ Choudhury S, Baradaran-Mashinchi P, Torres MP (May 2018). "Negative Feedback Phosphorylation of Gγ Subunit Ste18 and the Ste5 Scaffold Synergistically Regulates MAPK Activation in Yeast". Cell Reports. 23 (5): 1504–1515. doi:10.1016/j.celrep.2018.03.135. PMC 5987779. PMID 29719261.
- ^ Chen RE, Thorner J (August 2007). "Function and regulation in MAPK signaling pathways: lessons learned from the yeast Saccharomyces cerevisiae". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1773 (8): 1311–1340. doi:10.1016/j.bbamcr.2007.05.003. PMC 2031910. PMID 17604854.
- ^ Ariño J, Casamayor A, González A (January 2011). "Type 2C protein phosphatases in fungi". Eukaryotic Cell. 10 (1): 21–33. doi:10.1128/EC.00249-10. PMC 3019798. PMID 21076010.
- ^ Nagiec MJ, McCarter PC, Kelley JB, Dixit G, Elston TC, Dohlman HG (September 2015). Boone C (ed.). "Signal inhibition by a dynamically regulated pool of monophosphorylated MAPK". Molecular Biology of the Cell. 26 (18): 3359–3371. doi:10.1091/mbc.e15-01-0037. PMC 4569323. PMID 26179917.
- ^ Gartner A, Nasmyth K, Ammerer G (July 1992). "Signal transduction in Saccharomyces cerevisiae requires tyrosine and threonine phosphorylation of FUS3 and KSS1". Genes & Development. 6 (7): 1280–1292. doi:10.1101/gad.6.7.1280. PMID 1628831.
- ^ Martins BM, Swain PS (2013-08-08). Mac Gabhann F (ed.). "Ultrasensitivity in phosphorylation-dephosphorylation cycles with little substrate". PLOS Computational Biology. 9 (8): e1003175. Bibcode:2013PLSCB...9E3175M. doi:10.1371/journal.pcbi.1003175. PMC 3738489. PMID 23950701.
- ^ Ariño J, Velázquez D, Casamayor A (April 2019). "Ser/Thr protein phosphatases in fungi: structure, regulation and function". Microbial Cell. 6 (5): 217–256. doi:10.15698/mic2019.05.677. PMC 6506691. PMID 31114794.
- ^ Li Y, Roberts J, AkhavanAghdam Z, Hao N (December 2017). "Mitogen-activated protein kinase (MAPK) dynamics determine cell fate in the yeast mating response". The Journal of Biological Chemistry. 292 (50): 20354–20361. doi:10.1074/jbc.AC117.000548. PMC 5733576. PMID 29123025.
- ^ Errede B, Vered L, Ford E, Pena MI, Elston TC (September 2015). Boone C (ed.). "Pheromone-induced morphogenesis and gradient tracking are dependent on the MAPK Fus3 binding to Gα". Molecular Biology of the Cell. 26 (18): 3343–3358. doi:10.1091/mbc.e15-03-0176. PMC 4569322. PMID 26179918.
- ^ Wang X, Tian W, Banh BT, Statler BM, Liang J, Stone DE (November 2019). "Mating yeast cells use an intrinsic polarity site to assemble a pheromone-gradient tracking machine". The Journal of Cell Biology. 218 (11): 3730–3752. doi:10.1083/jcb.201901155. PMC 6829655. PMID 31570500.
- ^ Marini G, Nüske E, Leng W, Alberti S, Pigino G (June 2020). Bloom K (ed.). "Reorganization of budding yeast cytoplasm upon energy depletion". Molecular Biology of the Cell. 31 (12): 1232–1245. doi:10.1091/mbc.E20-02-0125. PMC 7353153. PMID 32293990.
- ^ Bardwell L (2005). "A walk-through of the yeast mating pheromone response pathway". Peptides. 26 (2): 339–350. doi:10.1016/j.peptides.2004.10.002. PMC 3017506. PMID 15690603.
- ^ Liu X, Bardwell L, Nie Q (April 2010). "A combination of multisite phosphorylation and substrate sequestration produces switchlike responses". Biophysical Journal. 98 (8): 1396–1407. Bibcode:2010BpJ....98.1396L. doi:10.1016/j.bpj.2009.12.4307. PMC 2856190. PMID 20409458.
- ^ Bhaduri S, Pryciak PM (October 2011). "Cyclin-specific docking motifs promote phosphorylation of yeast signaling proteins by G1/S Cdk complexes". Current Biology. 21 (19): 1615–1623. Bibcode:2011CBio...21.1615B. doi:10.1016/j.cub.2011.08.033. PMC 3196376. PMID 21945277.
- ^ Gonçalves-Sá J, Murray A (August 2011). "Asymmetry in sexual pheromones is not required for ascomycete mating". Current Biology. 21 (16): 1337–1346. Bibcode:2011CBio...21.1337G. doi:10.1016/j.cub.2011.06.054. PMC 3159855. PMID 21835624.
- ^ Banderas A, Koltai M, Anders A, Sourjik V (August 2016). "Sensory input attenuation allows predictive sexual response in yeast". Nature Communications. 7 (1): 12590. Bibcode:2016NatCo...712590B. doi:10.1038/ncomms12590. PMC 5007329. PMID 27557894.
- ^ Muller N, Piel M, Calvez V, Voituriez R, Gonçalves-Sá J, Guo CL, et al. (April 2016). Rao CV (ed.). "A Predictive Model for Yeast Cell Polarization in Pheromone Gradients". PLOS Computational Biology. 12 (4): e1004795. Bibcode:2016PLSCB..12E4795M. doi:10.1371/journal.pcbi.1004795. PMC 4831791. PMID 27077831.
- ^ Brenner S, Miller JH (2001). "Gene Rearrangement in Eukaryotic Organisms". Encyclopedia of Genetics. Academic Press. pp. 798–800. doi:10.1006/rwgn.2001.0518. ISBN 978-0-12-227080-2.
- ^ Coughlan AY, Lombardi L, Braun-Galleani S, Martos AA, Galeote V, Bigey F, et al. (April 2020). "The yeast mating-type switching endonuclease HO is a domesticated member of an unorthodox homing genetic element family". eLife. 9. doi:10.7554/eLife.55336. PMC 7282813. PMID 32338594.
- ^ Laney JD, Hochstrasser M (September 2003). "Ubiquitin-dependent degradation of the yeast Mat(alpha)2 repressor enables a switch in developmental state". Genes & Development. 17 (18): 2259–2270. doi:10.1101/gad.1115703. PMC 196463. PMID 12952895.
- ^ Wang Y, Lo WC, Chou CS (November 2017). Komarova NL (ed.). "A modeling study of budding yeast colony formation and its relationship to budding pattern and aging". PLOS Computational Biology. 13 (11): e1005843. Bibcode:2017PLSCB..13E5843W. doi:10.1371/journal.pcbi.1005843. PMC 5697893. PMID 29121651.
- ^ Liu D (2009). Handbook of nucleic acid purification. Boca Raton: CRC Press. p. 174. ISBN 9781420070972. OCLC 614294429.
- ^ Shore D (January 1997). "Genetic recombination: sex-change operations in yeast". Current Biology. 7 (1): R24–R27. Bibcode:1997CBio....7..R24S. doi:10.1016/S0960-9822(06)00012-1. PMID 9072164.
- ^ Klar AJ (October 2010). "The yeast mating-type switching mechanism: a memoir". Genetics. 186 (2): 443–449. doi:10.1534/genetics.110.122531. PMC 2942867. PMID 20940334.
- ^ Weber JM, Ehrenhofer-Murray AE (December 2010). "Design of a minimal silencer for the silent mating-type locus HML of Saccharomyces cerevisiae". Nucleic Acids Research. 38 (22): 7991–8000. doi:10.1093/nar/gkq689. PMC 3001064. PMID 20699276.
- ^ Maroc L, Zhou-Li Y, Boisnard S, Fairhead C (October 2020). Heitman J (ed.). "A single Ho-induced double-strand break at the MAT locus is lethal in Candida glabrata". PLOS Genetics. 16 (10): e1008627. doi:10.1371/journal.pgen.1008627. PMC 7591073. PMID 33057400.
- ^ Faure G, Jézéquel K, Roisné-Hamelin F, Bitard-Feildel T, Lamiable A, Marcand S, et al. (February 2019). Wolfe K (ed.). "Discovery and Evolution of New Domains in Yeast Heterochromatin Factor Sir4 and Its Partner Esc1". Genome Biology and Evolution. 11 (2): 572–585. doi:10.1093/gbe/evz010. PMC 6394760. PMID 30668669.
- ^ Thon G, Maki T, Haber JE, Iwasaki H (April 2019). "Mating-type switching by homology-directed recombinational repair: a matter of choice". Current Genetics. 65 (2): 351–362. doi:10.1007/s00294-018-0900-2. PMC 6420890. PMID 30382337.
- ^ Krebs JE, Kuo MH, Allis CD, Peterson CL (June 1999). "Cell cycle-regulated histone acetylation required for expression of the yeast HO gene". Genes & Development. 13 (11): 1412–1421. doi:10.1101/gad.13.11.1412. PMC 316758. PMID 10364158.
- ^ a b Butler G, Kenny C, Fagan A, Kurischko C, Gaillardin C, Wolfe KH (February 2004). "Evolution of the MAT locus and its Ho endonuclease in yeast species". Proceedings of the National Academy of Sciences of the United States of America. 101 (6): 1632–1637. Bibcode:2004PNAS..101.1632B. doi:10.1073/pnas.0304170101. PMC 341799. PMID 14745027.
- ^ Pâques F, Haber JE (June 1999). "Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae". Microbiology and Molecular Biology Reviews. 63 (2): 349–404. doi:10.1128/MMBR.63.2.349-404.1999. PMC 98970. PMID 10357855.
- ^ Belton JM, Lajoie BR, Audibert S, Cantaloube S, Lassadi I, Goiffon I, et al. (December 2015). "The Conformation of Yeast Chromosome III Is Mating Type Dependent and Controlled by the Recombination Enhancer". Cell Reports. 13 (9): 1855–1867. doi:10.1016/j.celrep.2015.10.063. PMC 4681004. PMID 26655901.
- ^ Kaplun L, Ivantsiv Y, Bakhrat A, Raveh D (December 2003). "DNA damage response-mediated degradation of Ho endonuclease via the ubiquitin system involves its nuclear export". The Journal of Biological Chemistry. 278 (49): 48727–48734. doi:10.1074/jbc.M308671200. PMID 14506225.
- ^ Houston P, Simon PJ, Broach JR (March 2004). "The Saccharomyces cerevisiae recombination enhancer biases recombination during interchromosomal mating-type switching but not in interchromosomal homologous recombination". Genetics. 166 (3): 1187–1197. doi:10.1534/genetics.166.3.1187. PMC 1470794. PMID 15082540.
- ^ Wu C, Weiss K, Yang C, Harris MA, Tye BK, Newlon CS, et al. (June 1998). "Mcm1 regulates donor preference controlled by the recombination enhancer in Saccharomyces mating-type switching". Genes & Development. 12 (11): 1726–1737. doi:10.1101/gad.12.11.1726. PMC 316872. PMID 9620858.
- ^ Ruan C, Workman JL, Simpson RT (October 2005). "The DNA repair protein yKu80 regulates the function of recombination enhancer during yeast mating type switching". Molecular and Cellular Biology. 25 (19): 8476–8485. doi:10.1128/MCB.25.19.8476-8485.2005. PMC 1265738. PMID 16166630.
- ^ Dodson AE, Rine J (November 2016). "Donor Preference Meets Heterochromatin: Moonlighting Activities of a Recombinational Enhancer in Saccharomyces cerevisiae". Genetics. 204 (3): 1065–1074. doi:10.1534/genetics.116.194696. PMC 5105842. PMID 27655944.
- ^ Eckert-Boulet N, Rothstein R, Lisby M (2011), Tsubouchi H (ed.), "Cell Biology of Homologous Recombination in Yeast", DNA Recombination, vol. 745, Totowa, NJ: Humana Press, pp. 523–536, doi:10.1007/978-1-61779-129-1_30, ISBN 978-1-61779-128-4, PMC 3968689, PMID 21660714
- ^ Ruderfer DM, Pratt SC, Seidel HS, Kruglyak L (September 2006). "Population genomic analysis of outcrossing and recombination in yeast". Nature Genetics. 38 (9): 1077–1081. doi:10.1038/ng1859. PMID 16892060. S2CID 783720.
- ^ Birdsell JA, Wills C (2003). "The evolutionary origin and maintenance of sexual recombination: A review of contemporary models.". In MacIntyre RJ, Clegg MT (eds.). Evolutionary Biology. Evolutionary Biology Series. Vol. 33. Springer. pp. 27–137. ISBN 978-0306472619.
- ^ Davey J (December 1998). "Fusion of a fission yeast". Yeast. 14 (16): 1529–1566. doi:10.1002/(SICI)1097-0061(199812)14:16<1529::AID-YEA357>3.0.CO;2-0. PMID 9885154. S2CID 44652765.
- ^ Bernstein C, Johns V (April 1989). "Sexual reproduction as a response to H2O2 damage in Schizosaccharomyces pombe". Journal of Bacteriology. 171 (4): 1893–1897. doi:10.1128/jb.171.4.1893-1897.1989. PMC 209837. PMID 2703462.
- ^ Staleva L, Hall A, Orlow SJ (December 2004). "Oxidative stress activates FUS1 and RLM1 transcription in the yeast Saccharomyces cerevisiae in an oxidant-dependent Manner". Molecular Biology of the Cell. 15 (12): 5574–5582. doi:10.1091/mbc.e04-02-0142. PMC 532035. PMID 15385622.
- ^ a b c Lin X, Hull CM, Heitman J (April 2005). "Sexual reproduction between partners of the same mating type in Cryptococcus neoformans". Nature. 434 (7036): 1017–1021. Bibcode:2005Natur.434.1017L. doi:10.1038/nature03448. PMID 15846346. S2CID 3195603.
- ^ Michod RE, Bernstein H, Nedelcu AM (May 2008). "Adaptive value of sex in microbial pathogens" (PDF). Infection, Genetics and Evolution. 8 (3): 267–285. Bibcode:2008InfGE...8..267M. doi:10.1016/j.meegid.2008.01.002. PMID 18295550.
Further reading
[edit]- Scott MP, Matsudaira P, Lodish H, Darnell J, Zipursky L, Kaiser CA, et al. (2004). Molecular Cell Biology (Fifth ed.). WH Freeman and Col, NY. ISBN 978-0-7167-4366-8.
- "Fus3". Saccharomyces Genome Database. SGD Project. Retrieved 21 March 2014.
